Forge Update: Multi-Omics Integration
I wrote for Metabolon about the value of multi-omics. Specifically, I drew parallels to Alexander von Humboldt’s conceptualization of the world as an ecosystem and how we have to measure it. It turns out that we didn’t solve everything with the Human Genome Project, and we’re still searching for ways to efficiently and effectively measure these incredibly dynamic, complex, and unobservable systems. A major focus, as I mentioned in that piece and have acted on here, is integrating multiple paradigms of large-scale data, particularly ‘omics data, to probe the entire industry of a cell.
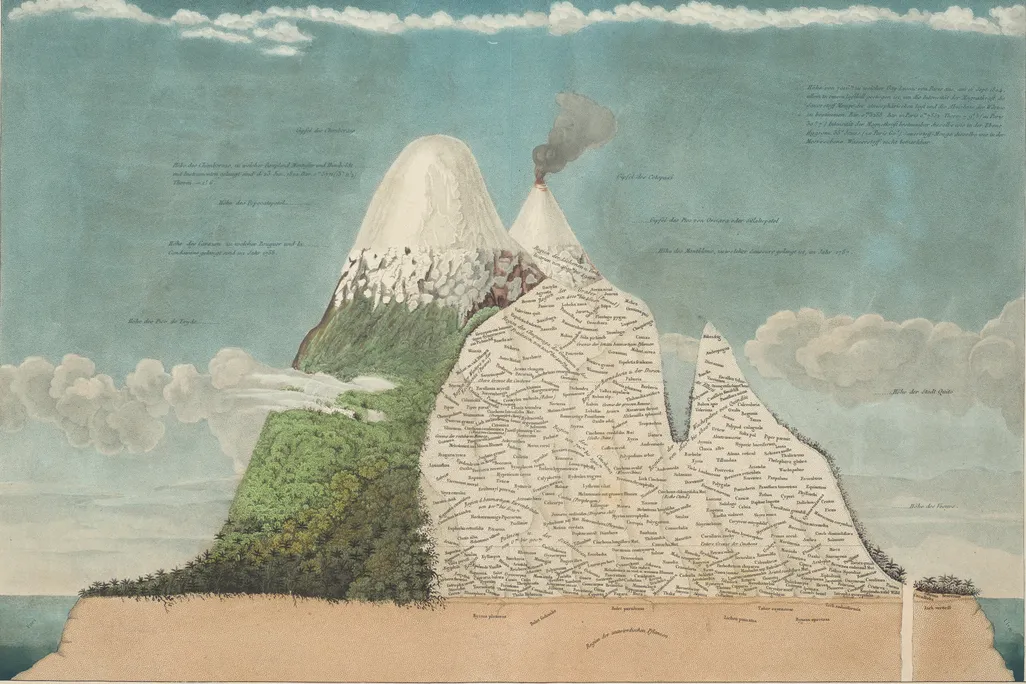

We’re in a weird but cool spot: instrument sensitivity is improving rapidly, costs are falling, and novel detection methods are being developed. We’re a long way from a multi-million dollar processing run of a single set of representative genomes: we can now design experiments that profile the small molecule chemistry of multiple sample types, the genome of the host, epigenetic regulation, and proteomic activity all at once, should we so choose (and if the funding allows). We can even apply that to larger sample sets to derive the throughlines between biomolecular forces and the phenotypes they provoke.
These changes have rebalanced the onus from sample acquisition/processing to experimental design/final analysis. It’s attractive to throw AI at this problem: isn’t it intended for large-scale data management and kitchen-sink analyses to derive biological hypotheses? This is where I’d exercise caution for several reasons:
- AI only has the context you give it. As scientists, I think Laplace’s Demon is a seductive way to reduce the universe. However, even with the tremendous advances we have in measuring and identifying biological phenomena, our brains are a lot more efficient at cataloguing purpose, context, and condition of the experiment.
- We’re still discovering. AI isn’t truly generative; it relies on the inputs of a vast number of experts and opinions to build its models. At this point, I’d hazard the opinion that we still don’t even know what we mean by “multi-omics”, much less AI. A lot of what’s going to become best practice in the future is predicated on our experimentation with the data afterward. We need iterative, guided approaches to learning before we can pass it off to an automated researcher.
- The data are most powerful in the hands of someone who understands the analytical process. We’re in a place where our labs and data acquisition protocols have been abstracted away from us and can be performed far away by other experts or machines. However, someone who can understand the statistical distribution of transcriptomics or the best statistical model for A/B testing in metabolomics (where absence is as much a signal as presence) is better suited to preprocess the data than an automated pipeline. This guidance is what stops us from having the same signals (looking at you, glutamine, IMPK, and HEXK) from drawing us back in, over and over.
A month ago I published the initial Forge post. At that point the platform handled single-assay preprocessing and analysis well; multi-omics integration was a labeled tab with a “coming soon” message. That tab now works.
Website is live here: www.insilijo.science. Registration costs one email (so I know who you are).
Dataset Management
Filtering
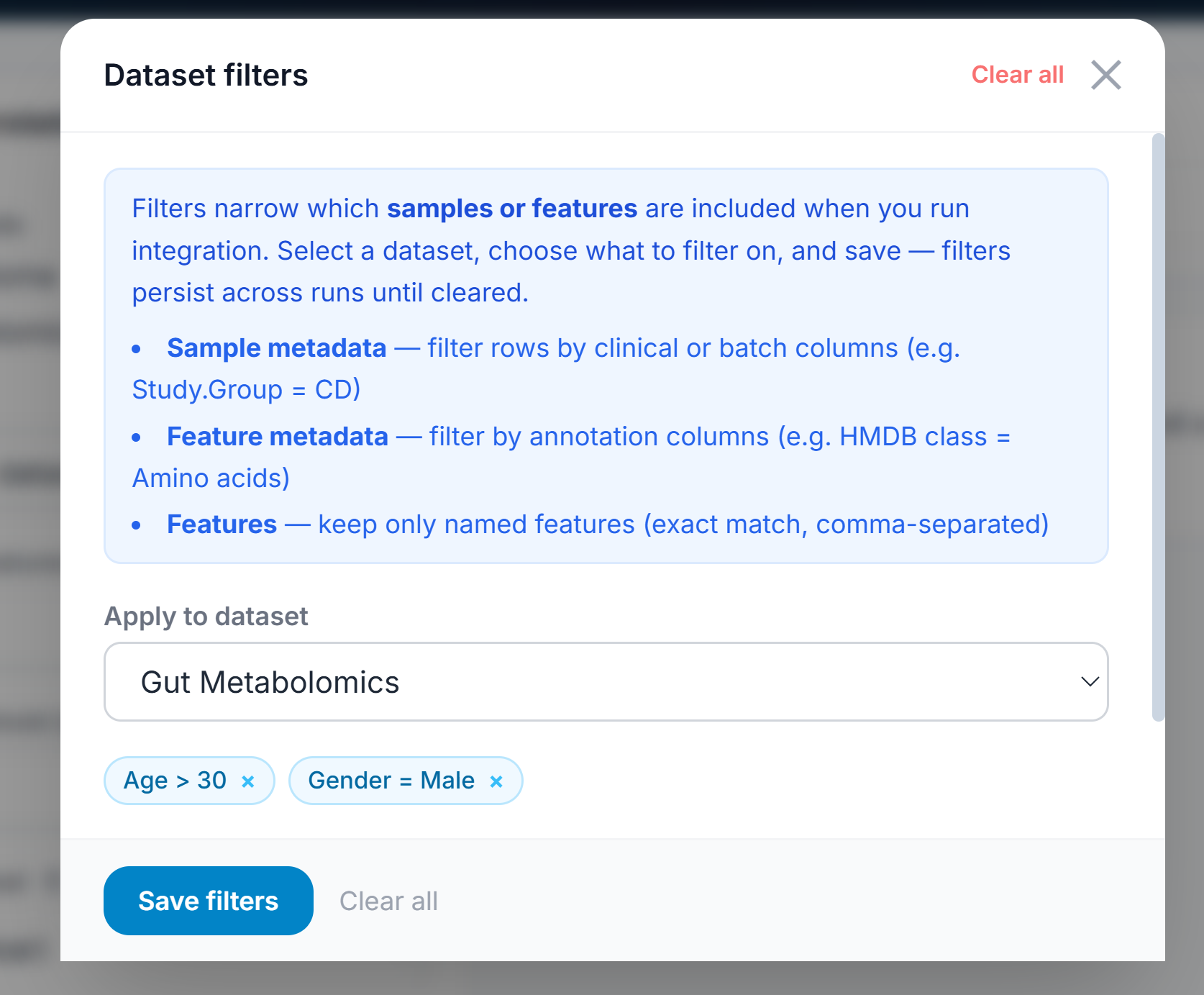
I’ve added filtering as an essential mechanism for the app. You can find that at the top right, and it will conditionally section off features or samples based on metadata or statistical quantities. That preset can be saved per user and generally applied. This allows a user to iteratively investigate intentional or unintentional dataset parameters. For example, you can filter out near-zero variant features and then isolate only patients with severe phenotype on PCA and do the same with moderate phenotype, giving a perspective on how the disease manifests at its most advanced. These filtered datasets can be duplicated as filtered versions to retain provenance but also segment the data intentionally.
Outlier Detection
Outliers can be detected either in single or multi-omic (concatenated) analysis. The user can flag them in the metadata, remove them from future analyses, or simply highlight them. Four methods are available: PCA + Mahalanobis Distance, Isolation Forest, Local Outlier Factor, and Robust MAD Z-score.
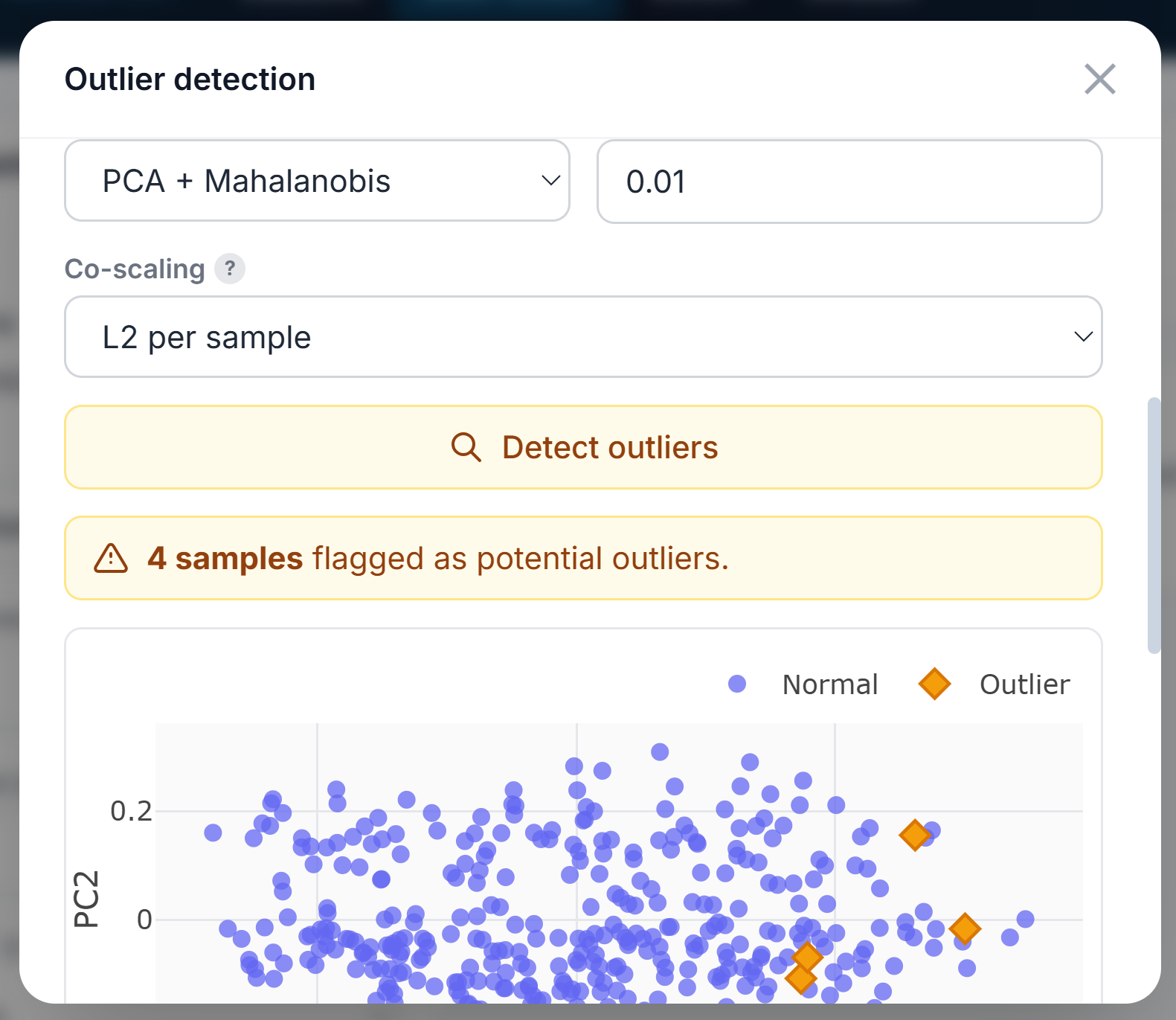
Managing
Derivative datasets (PCA, PLS-DA loadings, etc.) can be saved as alternative datasets. This is particularly valuable if you want to do any sort of modeling on loadings over time. I’m sure this’ll get more power as I we figure this out.
Multi-Omics Integration
The central addition is a full multi-omics analysis module backed by seven integration algorithms, each suited to a different study design and question. It’s designed to operate as close to single-dataset workflows as possible, since we’re hypothetically interrogating linked questions. One of my core philosophical positions on multi-omics is that no one really knows what it is. Is it the integration of orthogonal modalities? Do readouts of 10 lipids count as “metabolomics”? Can we slap timepoint 1 on timepoint 3 and call that multi-omics?
The core utility of this integration is to be flexible. Because multi-omics is flexible. I think a brilliant study design can be measuring the fecal microbiome, plasma metabolomics, and fecal metabolomics. Typically, different matrices with the same ‘omic wouldn’t qualify as “multi-omics”, but they’re answering different parts of the question. So have at it.
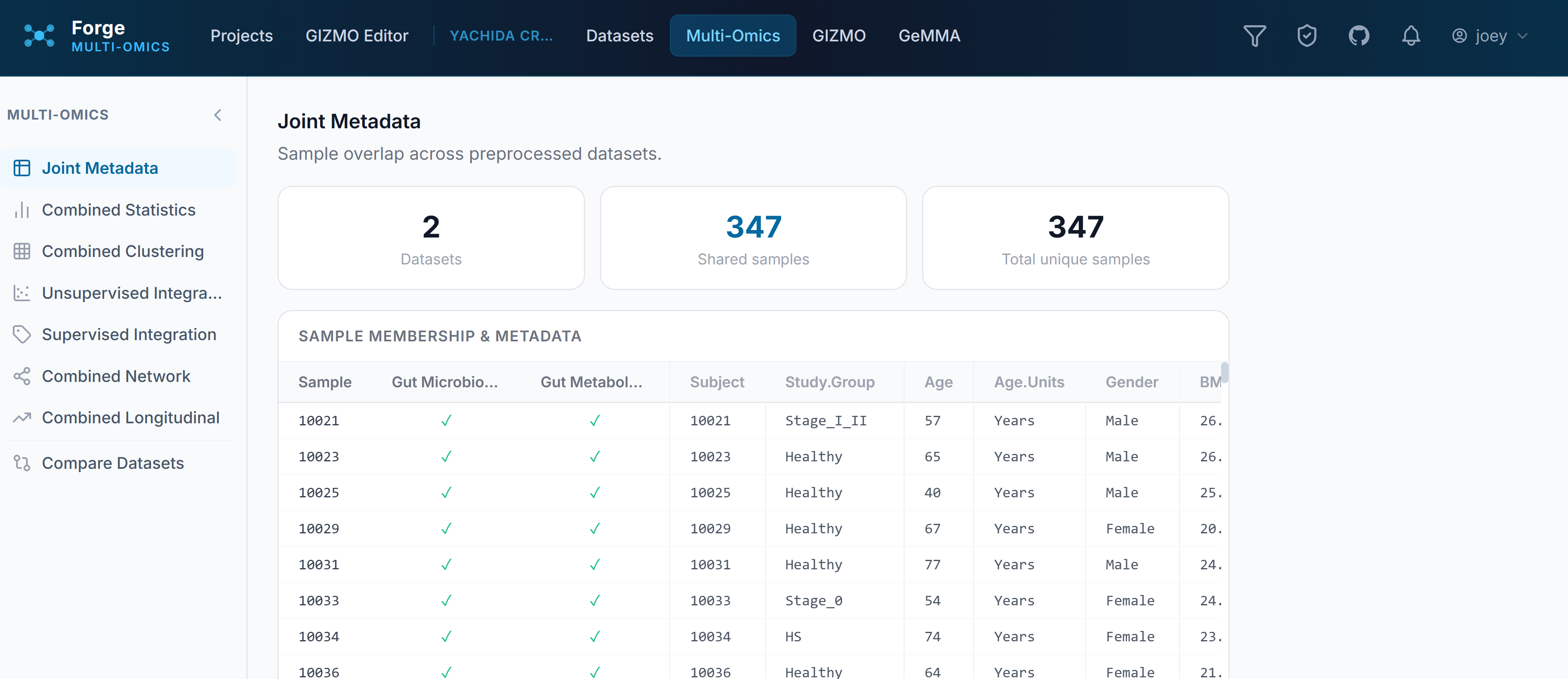
Joint Metadata
This shows where sample IDs overlap across datasets so you can verify that the metadata you need for grouping carries through the merge. By default, shared sample names drive horizontal concatenation to retain total sample metadata — so you’ll see things like “Read Depth” applied to a metabolomics dataset because it’s inheriting from the metagenomic data. If sample IDs aren’t lining up, this is where to catch it before going further.
Combined Statistics
We have PCA. We have PLS-DA. We love them both. No need to dispense with your heavy hitters. The value here – and generally – is that you can preprocess and re-scale for each application. This means that we can directly determine the best scaling routine to match our QC expectations (e.g., batch number isn’t the primary driver of group separation) or best resolution in expectation (healthy is substantially different than diseased on PC1).
Unsupervised Integration
MCIA (Multi-Co-Inertia Analysis) finds a common ordination across multiple omics layers simultaneously: useful when you want to visualize sample structure across assays without a supervised target. Samples that cluster together in metabolomics should cluster together in transcriptomics if the biology is coherent; MCIA makes that concordance (or its absence) visible.
JIVE (Joint and Individual Variation Explained) decomposes the variance in each omics layer into a joint component shared across assays and individual components unique to each. This is valuable when you expect some signal to be shared (a treatment effect visible in both metabolomics and proteomics) and some to be assay-specific (technical variation, platform-specific features). JIVE separates those contributions explicitly.
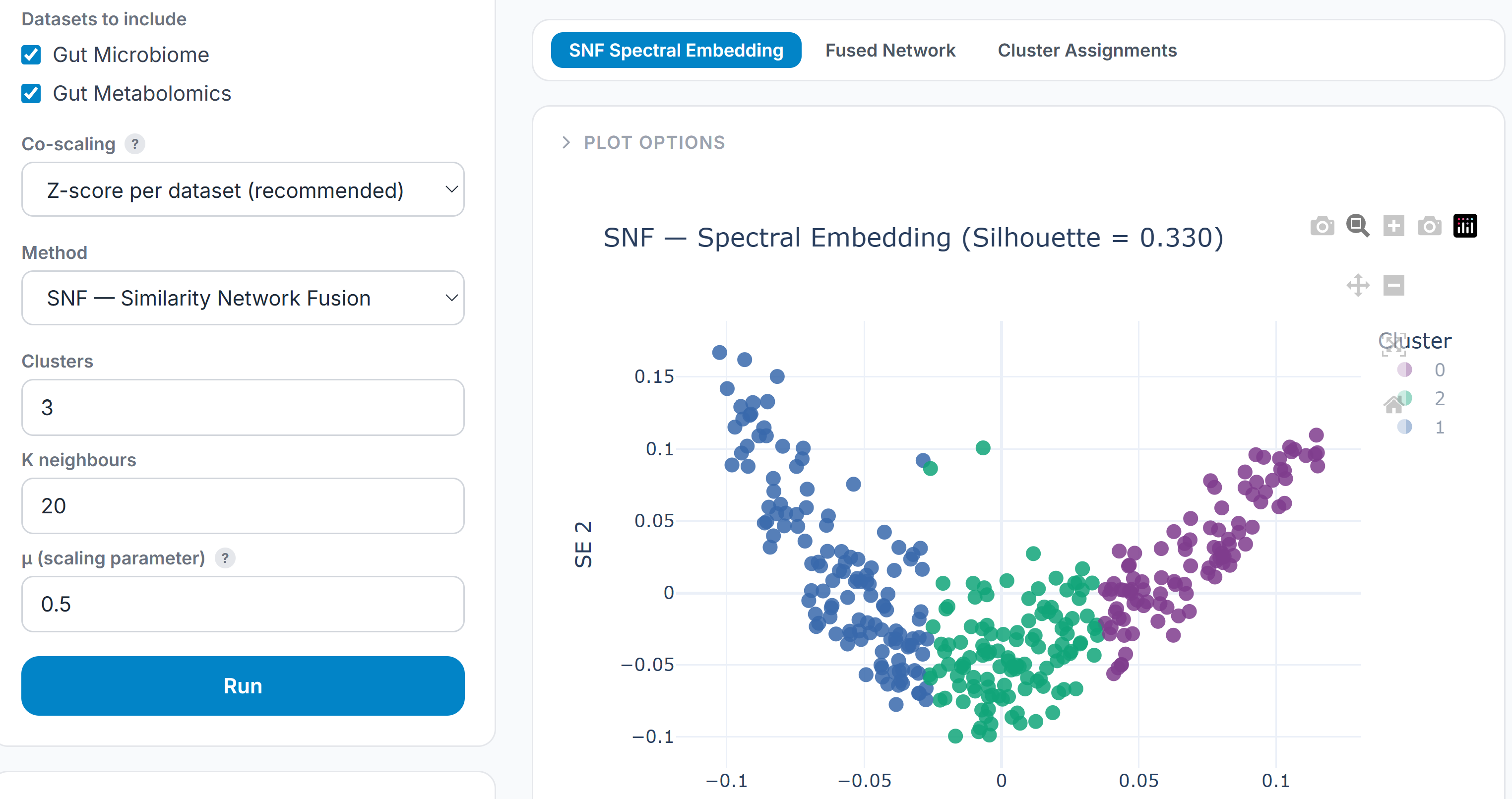
SNF (Similarity Network Fusion) constructs a similarity network for each omics layer separately, then iteratively fuses them into a single patient similarity network. SNF is particularly suited to subtype discovery: it finds patient groupings that are consistent across assays without requiring that the same features drive the separation in each one.
MOFA+ is a probabilistic latent factor model that identifies axes of variation shared across omics layers. Unlike JIVE, which is deterministic, MOFA+ provides uncertainty estimates and handles missing values natively, making it well-suited to datasets where not every sample has every assay.
Variance Partitioning estimates how much of the variance in each feature is attributable to each omics layer, to covariates, and to their interactions. The output is a feature-level decomposition that identifies which metabolites are primarily driven by transcriptional regulation, which by microbiome activity, which by clinical covariates — a direct answer to the question of what is actually explaining the signal.
Supervised Integration
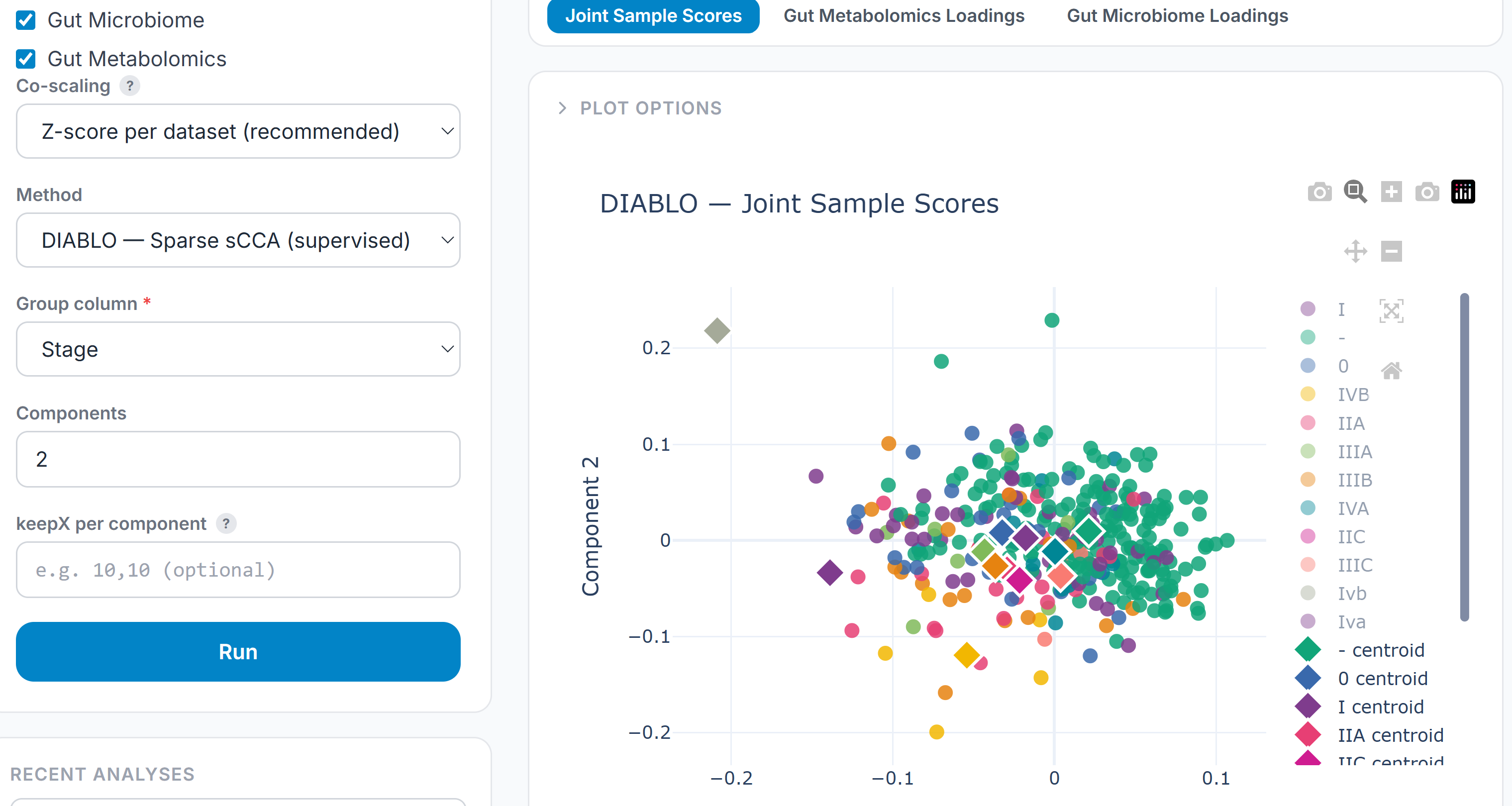
DIABLO is the supervised counterpart to MCIA — a sparse canonical correlation approach that finds the linear combinations of features across assays that best discriminate between groups. Where MCIA asks “what structure is shared?”, DIABLO asks “which features, across which assays, best explain this outcome?”
Random Forest Late Fusion trains independent models on each omics layer, then combines the predictions using a second-stage ensemble. This is the pragmatic approach when the assays are high-dimensional and sparse — it doesn’t require feature alignment across layers, tolerates missing data gracefully, and produces feature importances per layer that are directly interpretable.
The integration hub gates availability on dataset composition: MCIA and JIVE require at least two aligned assays; DIABLO and variance partitioning additionally require a group label. The interface surfaces which datasets are eligible for each method and why, so the user understands what the algorithm needs rather than hitting an error.
Combined Network
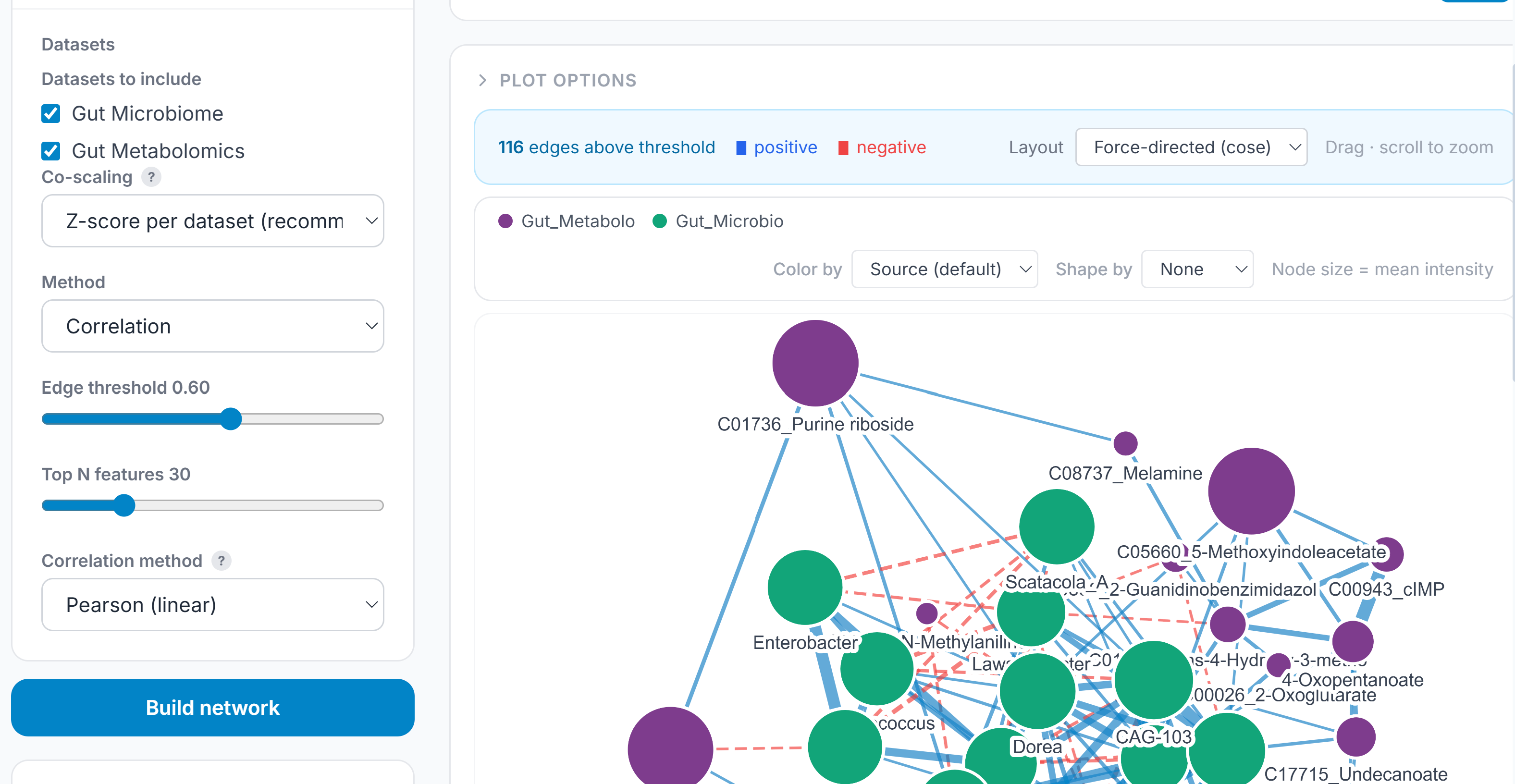
Single-assay correlation networks are useful; multi-omics correlation networks ask the harder question: which features co-vary not just within an assay but across them? The combined network builds from the aligned, concatenated multi-omics matrix and offers three modes depending on what the data and question support.
Pearson correlation constructs edges between all feature pairs using BH-corrected Fisher z-transform p-values across the full candidate set before any correlation threshold is applied — the same approach used in the single-assay network, now operating across assay boundaries. A metabolite–transcript edge carries the same statistical treatment as a metabolite–metabolite edge.
Partial correlation (Graphical Lasso) estimates conditional independence rather than marginal correlation. An edge between two features in a partial correlation network means they are associated after accounting for everything else — it removes the transitive connections that inflate standard correlation networks and leaves only the direct relationships. The regularization parameter is selected by cross-validation.
Spearman correlation ranks each feature’s values across samples before computing the correlation coefficient. This makes it robust to outliers and appropriate for data that is not normally distributed — which describes most metabolomics and proteomics features after any real preprocessing pipeline. Edges are constructed using the same Fisher z-transform significance testing as Pearson, applied to the rank correlation coefficients.
Kendall correlation also operates on ranks but uses concordant and discordant pair counts rather than a linear approximation. It is more conservative than Spearman — it produces lower correlation values for the same underlying relationship — but is more reliable at small sample sizes where Spearman’s approximation is less stable. Prefer Kendall when sample counts are low (roughly n < 30) or when the data has many tied ranks.
Compare Datasets
The dataset comparison utility operates on completed analytical results rather than raw data. Select any two datasets in a project; Forge extracts per-feature numeric vectors from their respective preprocessing runs and downstream analyses — fold change, VIP scores, mean intensity, coefficient of variation, network degree centrality, longitudinal slope — then compares them on shared features.
The core comparison is a scatter of matched feature values with Pearson and Spearman correlations reported. But the more useful output is the preprocessing alignment check: if two datasets have been normalized differently, scaled differently, or log-transformed with different bases, a high feature-level correlation may still be analytically invalid. Forge surfaces these preprocessing mismatches explicitly alongside the correlation, so the comparison is interpreted with the right caveats attached.
The utility also reports Jaccard overlap of the feature sets, shared grouping variables across analyses, and which analysis types are present in both datasets — the structural prerequisites for a valid comparison. The intent is to support replication analysis, batch experiment reconciliation, and cross-cohort validation without requiring the analyst to manually reconstruct the comparison from separate exports.
GIZMO, the Knowledge Graph
GIZMO, the underlying knowledge graph and my dearly departed, wheelchair-bound pug, has undergone a facelift. Edges now incorporate HMDB IDs and better connect to existing databases. I’ll be working to manually curate these edges in the background. It currently draws from 17 public databases:
| Database | Provides |
|---|---|
| Reactome | Reactions, pathways, EC numbers, gene associations |
| ChEBI | Compound structures, InChI, SMILES, mass, charge |
| MetaNetX | Cross-reference hub — ChEBI ↔ HMDB ↔ Reactome ↔ PubChem IDs, stoichiometry |
| MONDO | Disease ontology, IEM subset, ICD-10/OMIM/Orphanet cross-references |
| Open Targets | Scored gene–disease associations (GWAS, rare variant, expression, somatic) |
| Orphanet | Rare disease classifications, gene–disease associations |
| HPO | Phenotype terms, phenotype–gene and phenotype–disease associations |
| NCBI Gene | Gene identifiers, symbols, orthologs across human and 7 model organisms |
| GTEx | Tissue-specific expression across 54 tissues (median TPM) |
| STRING | Protein–protein interactions with confidence scores |
| ChEMBL | Drugs, targets, mechanisms of action, ADMET, toxicology assays |
| PubChem | Chemical properties, name→InChIKey resolution |
| EPA CompTox | Hazard flags (carcinogen, reproductive, neurotox), CAS numbers, DTXSID |
| CTD | Chemical–gene interactions, chemical–disease associations |
| T3DB | ~3,700 toxin records — targets, LD50, exposure routes |
| ClinVar | Pathogenic/likely pathogenic variants, clinical significance |
| Metabolon | 5,395 measured metabolites with platform metadata (PMC open-access subset) |
From here you can run enrichment analysis, causal chain queries, or druggability targeting. A feature mapper checks that your dataset’s identifiers align with the graph before any query runs. It supports multiple datasets simultaneously and accepts VIP scores, fold changes, or mean intensities as the input signal — whatever came out of your analysis. This is where I’ll put the most work, both with managing the integrations and creating hypothesis-driving options like:
- filtering the graph for your question,
- compartments,
- edge weighting,
- algorithms,
- more databases,
- applying GIZMO to statistical visualizations.
What’s Next
The immediate priorities are the end-to-end case study on a public dataset (MTBLS or Metabolomics Workbench), pathway enrichment visualization, and the software paper. The multi-omics module is functional but the visualization layer for cross-assay results is still being extended — the algorithms run, but the output deserves richer interactive plots than what currently ships.
GeMMA and GIZMO, the companion tools in the Insilijo stack, are each close to their own publication milestones. I’ll write about those separately as they clear validation.
If you are working with multi-omics data and want to talk about what Forge can do for your specific study design, reach out via the contact page or open an issue on GitHub.
Enjoy Reading This Article?
Here are some more articles you might like to read next: