Forge: A Multiomics Analytical Platform
Biology’s really hard. It’s hard to measure, it’s impossible to predict, and those long-held skeletal rules we believe about it – the central dogma, Mendelian genetics, what goes in must come out – are limited abstractions at best. It’s also incredibly necessary to understand the full impact of that heterogeneity: internal regulation, genetic differences, the accumulation of substrates, and many other outside-the-blueprint biological phenomena are what actually inform why diseases might occur, where people might react to different medication, and even how we might engineer bacteria to suitably fit into artificial scenarios to benefit the planet.

I have been building computational platforms for multiomics since graduate school, integrating linear optimization, transport phenomena, and canonical knowledge to interrogate heterogeneous and emergent behavior in cellular populations. This is convenient and slightly bibliographic: given what we know, what can we understand about what we cannot easily measure? It’s an interrogation of the indicative present vs. subjunctive; what could we know based on what we do know. George E.P. Box’s famous statistics quote gets thrown around here a lot: “All models are wrong, but some are useful.”
But how did we even get to the point where we could model?
At the core of all canonical models is a series of experiments that survive and are codified into understanding. With the advent of biobanks, big data, and AI, we’re running a lot of experiments. But those experiments don’t passively result in biological understanding. Data arrives as a bundled mess, in formats that do not agree with each other, where preprocessing decisions are made invisibly and inconsistently, represented as statistical results reported without the context needed to interpret them, and with an analytical archaeology held together by a combination of R scripts and institutional memory. Having more data, more AI, more hardware hasn’t solved this; it’s amplified it: every operation is more immediate and larger scale, but every choice is larger and more legible in the final product. In that paradigm, every organization is making its own Library of Babel at higher scale and much more rapidly; each siloed in its own way, each segmented and operationalized separately, and now equipped with a very willing robot partner to accelerate both the good and the bad.
My work is in that funnel: a process that takes new, multimodal data and refines it into something that fits within canonical systems or extends them reliably. The struggle is using the benefits of the signal while being conscious of the amplified noise, especially in multiomic contexts where multiple measurements stuck together from the same system can elucidate those complex biological features we need to understand.
Forge is the current iteration of that work. It is a production-deployed multiomics analytical platform, built and maintained under Insilijo Science, my independent consulting and advisory practice. This post is an introduction to what it does and why certain design decisions were made the way they were.
The Problem
Omics studies generate data at a scale and complexity that strain conventional analytical workflows. For example, a typical untargeted metabolomics experiment produces tens of thousands of features – many uncharacterized – distributed across samples with varying missingness, batch effects, and normalization requirements. The most common metabolomics data, which is widely targeted, still results in over a thousand features. On the other hand, transcriptomic studies can easily have tens of thousands of features with several thousand of them being differential. Affinity-based proteomics experiments are typically in the multiple thousands range. Getting from these raw feature matrices to a biologically interpretable result requires a sequence of decisions, each of which shapes the downstream analysis: which imputation strategy fits the missing data mechanism, whether normalization should be sample-level or feature-level or both, how aggressively to correct for run-order drift, which statistical framework fits the study design, and how should scaling be applied for non-normal distributions.
These decisions are not interchangeable; they interact. But in most workflows, they are made once, in an R or Python session, with no systematic record of the configuration, no interface for a collaborator to inspect or modify them, and no mechanism for reproducing the result on a different dataset without manual reconstruction. The upstream and downstream processes are decoupled and uncollaborative meaning that analyses inherit decisions that might not be appropriate for their interpretation. Transparency is not only nice, it is necessary: you can’t build an effective scientific argument based on decisions you don’t know about or don’t understand.
Forge is built around the premise that the analytical pipeline itself is a scientific object, something that should be configurable, documented, persistent, comprehensible, and shareable in the same way the data is. Science is a collaborative ecosystem, and the decisions embedded in data analysis must be as portable and composable as the data itself. Lastly, it should be low barrier-to-entry; a first year undergraduate student studying biology should be able to run a PCA for the first time and understand why, while a seasoned clinician shouldn’t have to learn Python to see expected group separation in A/B testing.
What Forge Does
Data Ingestion
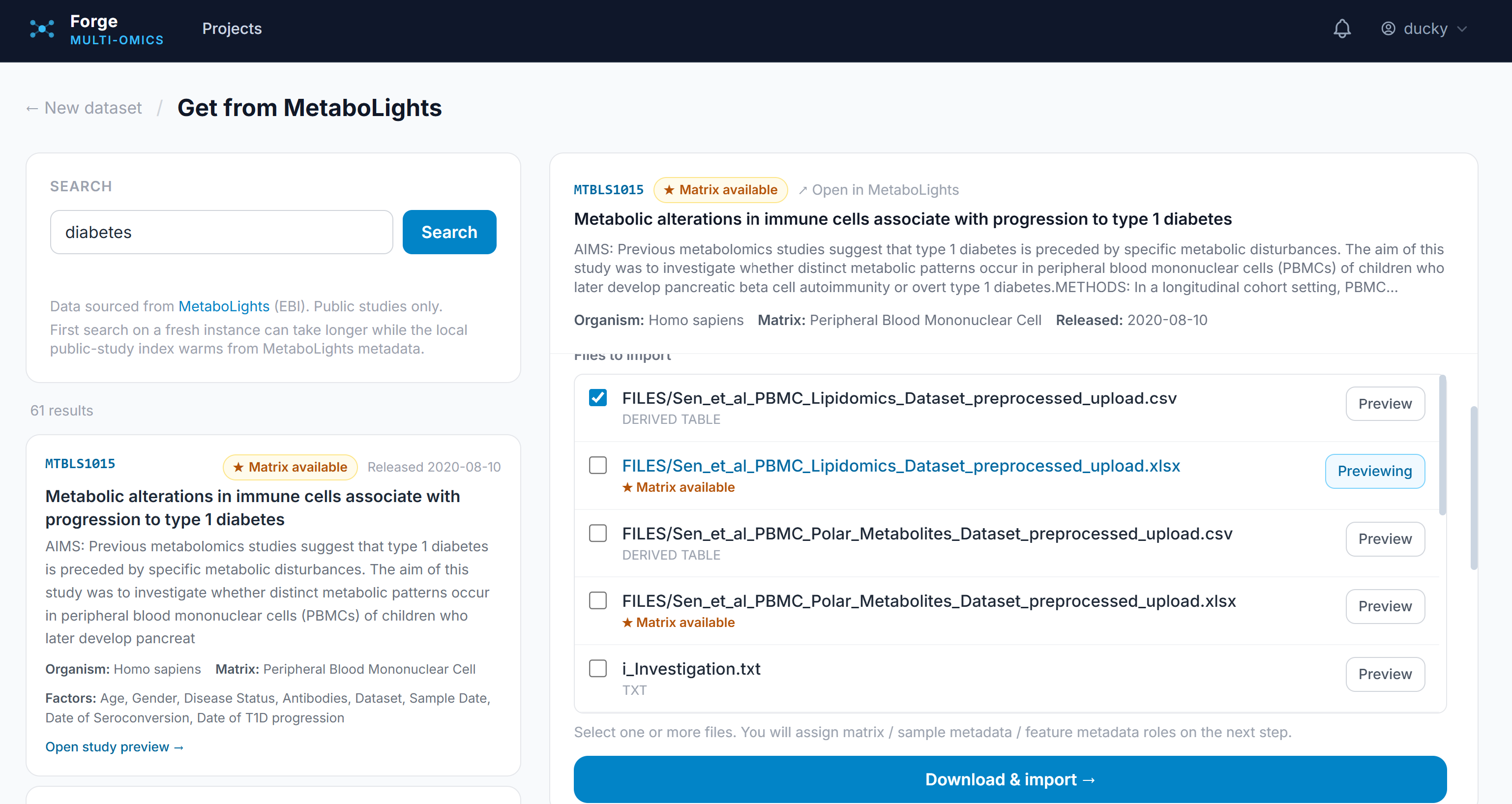
Forge accepts all tabular-style data outputs with integrations for four major data repositories: Metabolights and Metabolomics Workbench for metabolomics, PRIDE for proteomics, and GEO for gene-based experiments. It allows uploading of tabular data with a specification of the data type and currently contains 9 synthetic and 14 public datasets for exploration. Users, once registered, can create projects with multiple datasets to stay organized.
QC and Preprocessing Pipeline
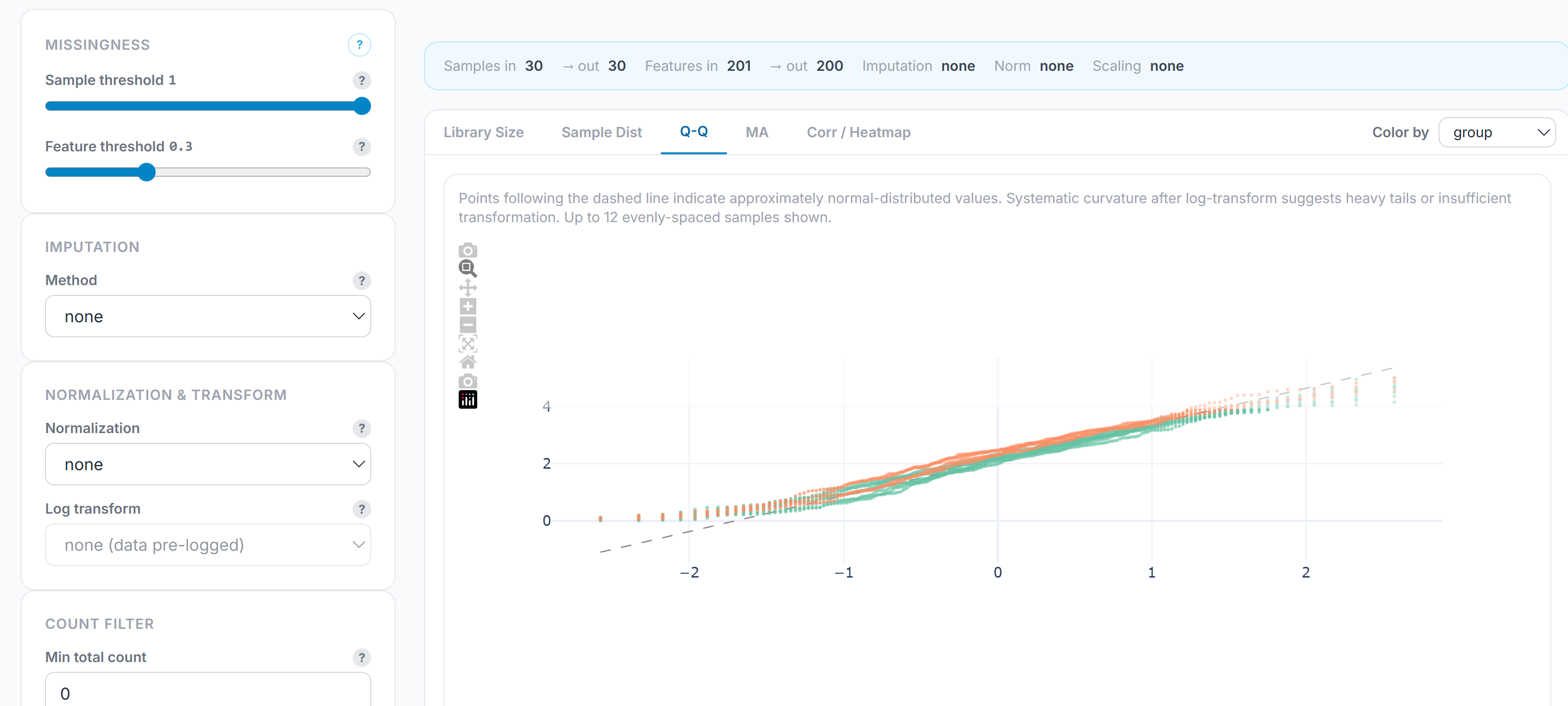
The preprocessing pipeline is a configurable sequence of steps exposed as a live control panel, specific for the data type. For example, for metabolomics, it enables missingness filtering, imputation (none, mean, median, half-minimum, KNN), normalization (total-area, median, quantile, internal standard, total protein), log transformation, scaling (standard, Pareto, min-max, robust), and batch correction. Similarly, specific pipelines exist for metagenomics, proteomics, and transcriptomics. Each parameter is a labeled control with a documented effect. Configurations are saved as named profiles and can be reloaded across datasets, which means a core facility or team can enforce consistent preprocessing across a study without re-specifying the pipeline by hand each time. Finally, QC plots – run order, TIC/MIC, Q/Q plots, correlation – are directly visible, enabling evaluation of analysis performance based on metadata, run order, and other configurable parameters.
Statistical Analysis
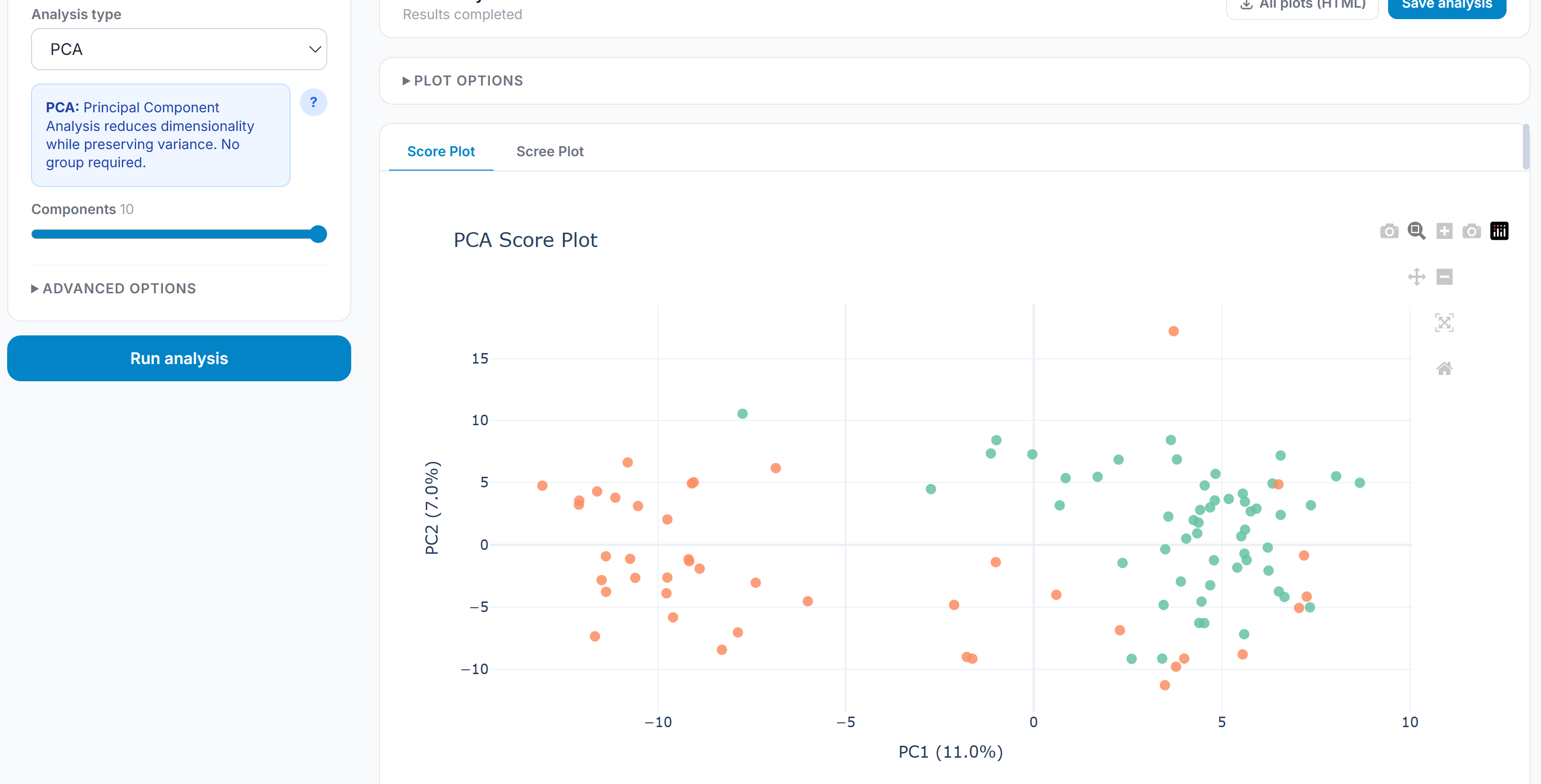
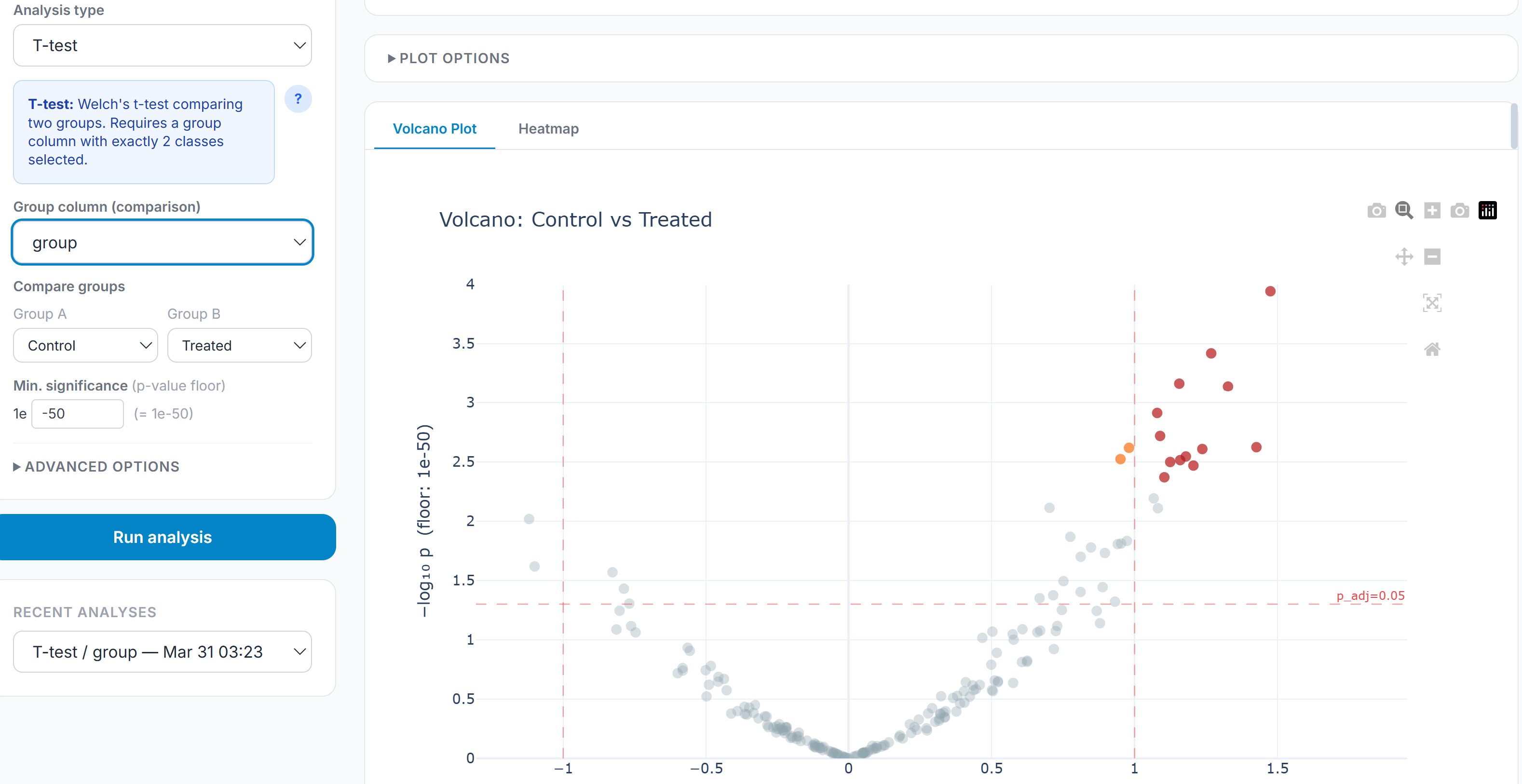
Differential abundance testing covers t-tests (Welch’s, Student’s, Mann-Whitney U), one-way ANOVA, and ANCOVA with covariate support. Effect sizes are reported alongside p-values for every test: Cohen’s d for pairwise comparisons, η² for ANOVA, R² for ANCOVA. Multiple testing correction is applied globally across all comparisons in an analysis using BH FDR, Bonferroni, or Holm-Šídák. PCA and PLS-DA handle dimensionality reduction and group separation; clicking any point on the scatter opens a feature contribution panel decomposing the score by loading per axis.
Longitudinal Analysis
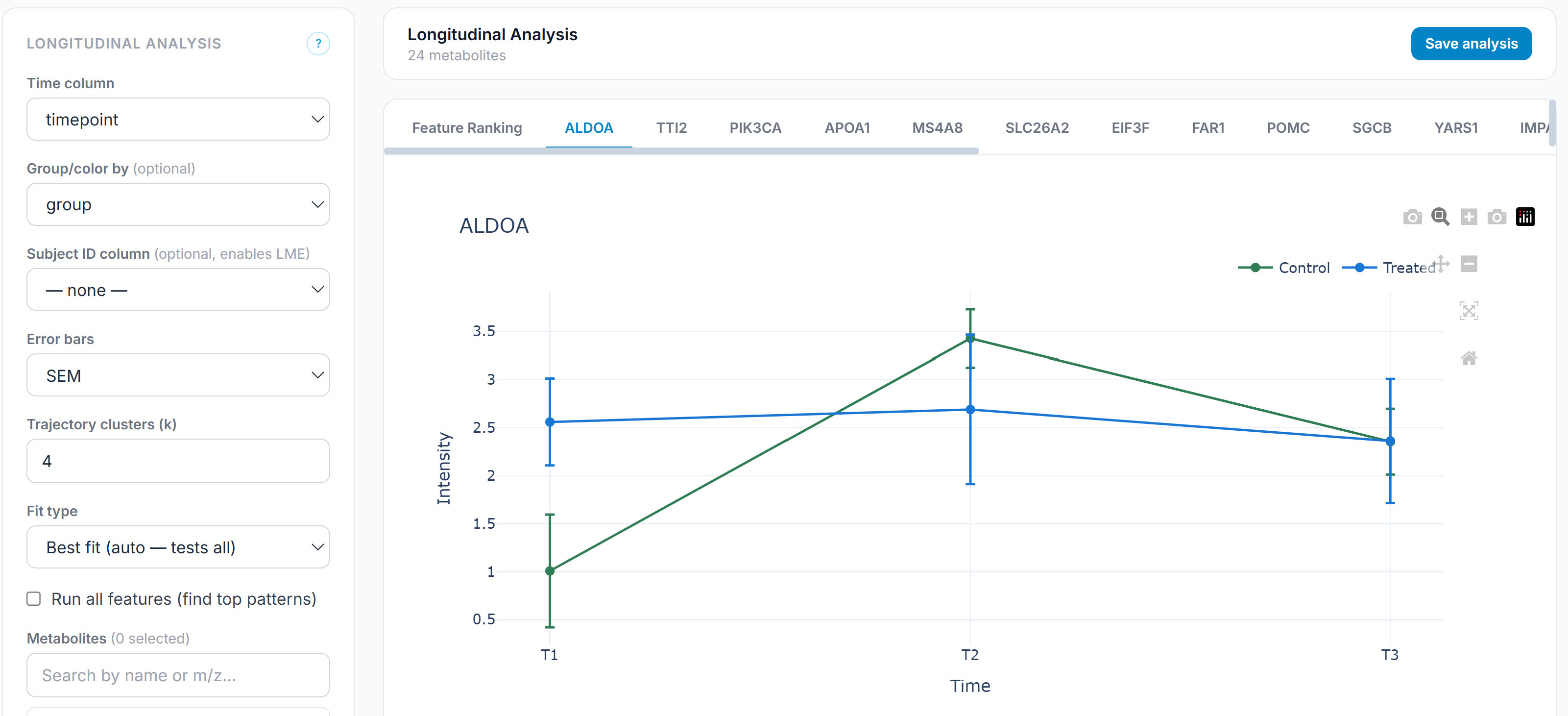
Support for longitudinal study designs is underrepresented in most analytical platforms, which are built around cross-sectional comparisons. Forge includes a dedicated longitudinal module that tracks feature trajectories across time points, models within-subject change, and identifies features with statistically significant time-course patterns. This matters for intervention studies, clinical trials, and any design where the relevant signal is the shape of change rather than a single-timepoint difference. The module also scans the full feature set and selects the best-fit trajectory model (time, space, or other continuous parameter) and reports the best model (linear, quadratic, logistic, or other) for that feature, enabling investigation of alternative longitudinal responses.
Correlation Network Visualization
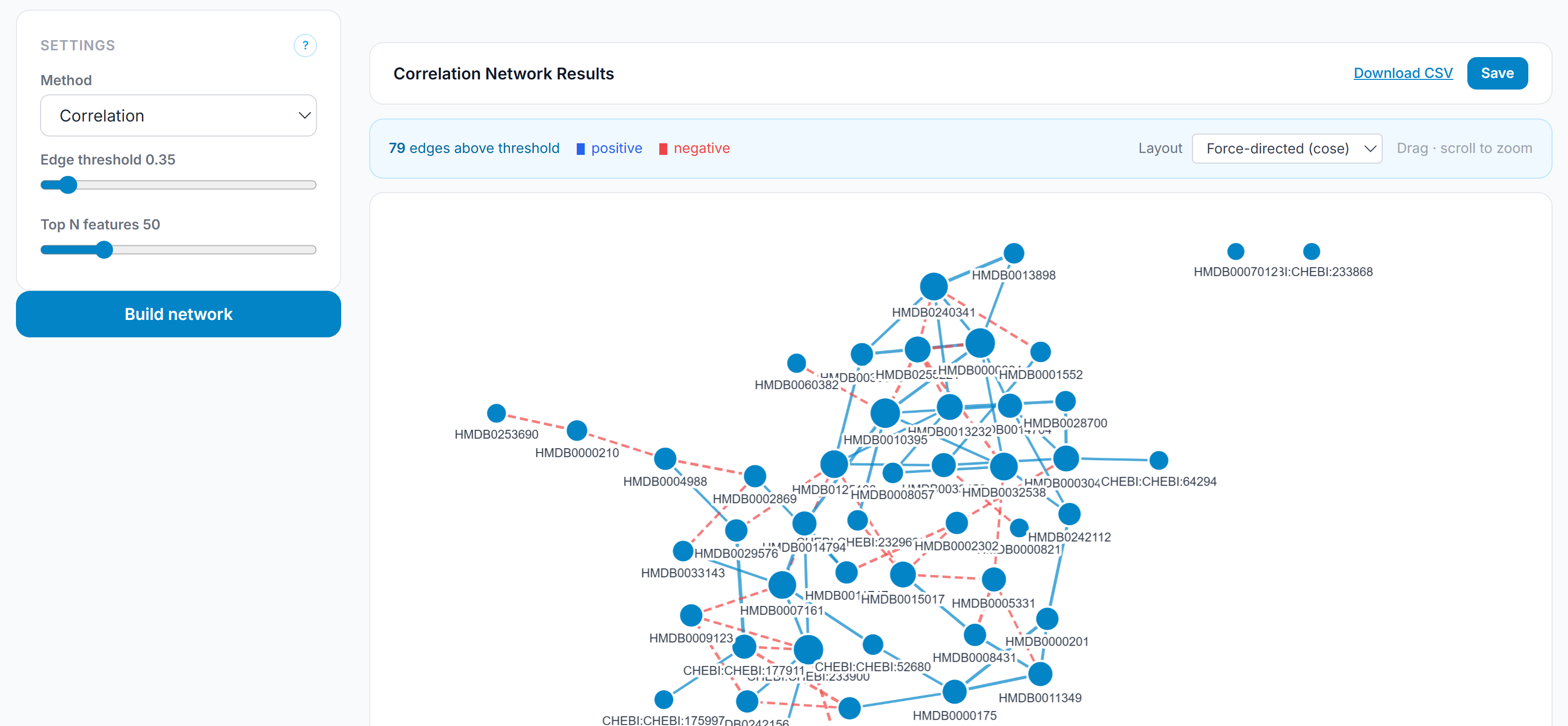
Standard metabolomics statistics are feature-centric: they ask which individual metabolites are different between conditions. Network analysis asks a different question — which features move together, and what does the structure of those co-variation relationships reveal about the underlying biology? Forge builds correlation networks from the preprocessed feature matrix, applies Fisher z-transform significance testing with BH correction on all candidate edges before threshold filtering, and renders the result as an interactive graph. This is a systems-level interpretive layer that sits on top of the standard statistical output, not a replacement for it.
Knowledge Graph and Pathway Context
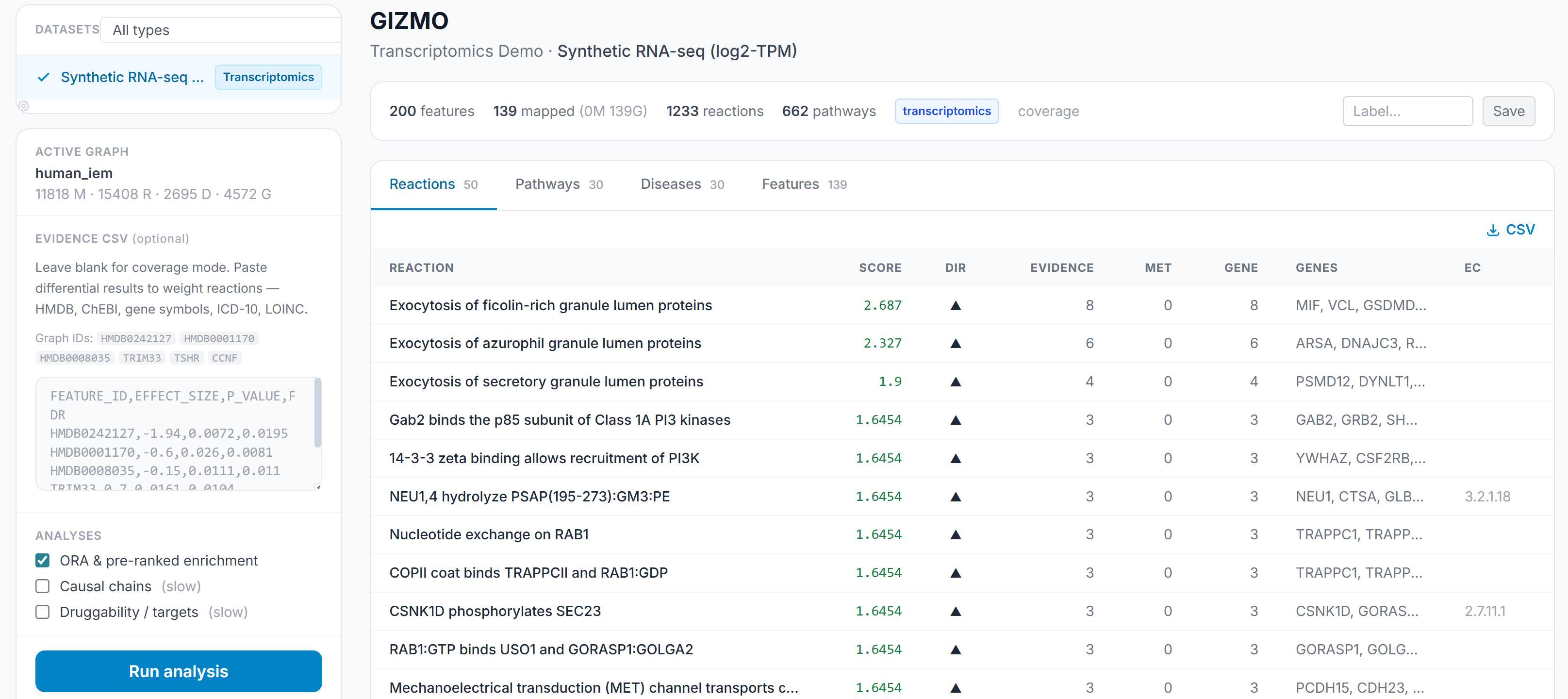
Forge integrates pathway context through a knowledge graph layer that maps features to biological pathways and provides enrichment-level interpretation alongside the feature-level statistics. This is where multiomics becomes interpretable to a biologist: not “these 47 features are significant” but “these features converge on these pathways, with this evidence, and with these other supporting features.” This also solves the ontology issue; teams collaborate at the biological, not the analytical, level. By integrating multiple open-source, public databases organizing different omics layers, phenotypes, and clinical data, Forge maintains a computation-ready knowledgegraph that can be directly applied to different contexts.
Insilijo Science
Forge is the analytical infrastructure I use in my consulting practice. Insilijo Science provides multiomics analysis, pipeline development, and data strategy advisory to life sciences teams: biotech, pharma, and academic groups working on metabolomics, proteomics, and integrated omics studies. If you are building a multiomics analytical capability and want to talk about what that looks like in practice, reach out.
What’s Next
The immediate roadmap includes pathway-centric visualization – enrichment drilldown and canonical pathway overlays rather than just feature-level output – and multi-omics integration proper: cross-assay sample alignment, late-fusion approaches, and coordinated visualization across omics layers. The knowledge graph integration will deepen as the companion tools in the Insilijo stack mature. Several software papers are in preparation for Forge and several other tools I’ve been working on.
The longer arc is the one I have been working on since graduate school: making the full path from measurement to biological meaning something that a research team can traverse reproducibly, collaboratively, and without losing the scientific accountability for every analytical decision along the way. Forge is the current best version of that.
Enjoy Reading This Article?
Here are some more articles you might like to read next: